Sangjin Kim
Probing the Microscopic Structure of a Cell from Diffusivity of the Proteome
 Cells are Nature’s test tubes, in which the biochemical reactions of life take place. Unlike laboratory test tubes, in which scientists can create a well-defined reaction condition, the physico-chemical properties of the cellular interior remain poorly understood. Recently, the importance of understanding the cellular interior has been increasingly emphasized due to novel findings from interdisciplinary research such as studies of liquid-liquid phase-separation inside a cell and efforts to engineer synthetic cells. A new technological breakthrough is urgently needed to characterize the cellular interior in greater detail.
Cells are Nature’s test tubes, in which the biochemical reactions of life take place. Unlike laboratory test tubes, in which scientists can create a well-defined reaction condition, the physico-chemical properties of the cellular interior remain poorly understood. Recently, the importance of understanding the cellular interior has been increasingly emphasized due to novel findings from interdisciplinary research such as studies of liquid-liquid phase-separation inside a cell and efforts to engineer synthetic cells. A new technological breakthrough is urgently needed to characterize the cellular interior in greater detail.
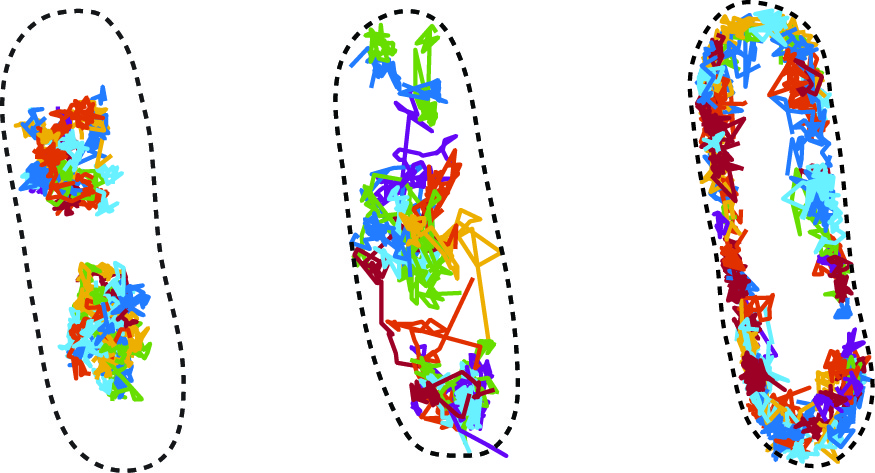
Professor Kim’s long-term research goal is to use physics-inspired approaches to determine the microscopic structure of cells. Toward this goal, her group is developing new technology for mapping protein-protein interactions and macromolecular crowding at the molecular scale in live bacterial cells. The innovation is based on precisely measuring the “jiggling” thermal motion of proteins that is as small as ~10 nanometers in 1-10 milliseconds. This microscopic dynamic reflects spatial details about the molecule’s immediate surroundings, hence offering an opportunity to profile macromolecules in the vicinity. The technology is based on single-particle tracking of proteins using single-molecule fluorescence microscopy and a machine learning-based analysis pipeline. This technology will enable scientists across multiple fields to obtain the high-resolution molecular picture of a cell’s interior and provide mechanistic and systems-level insights into how cells operate properly.
